Note
Go to the end to download the full example code.
GEDAI spectral🔗
This tutorial demonstrates how to use spectral GEDAI.
Spectral GEDAI is a frequency-specific denoising method that extends the
generalized eigenvalue decomposition approach of GEDAI.
Its approach focuses on isolating and removing artifacts within specific
frequency bands. For that, the spectral GEDAI first decomposes the EEG
data into its frequency components using wavelet transform, then applies
GEDAI to each frequency band separately. Finally, the denoised frequency
components are recombined to reconstruct the cleaned EEG signal.
import matplotlib.pyplot as plt
from mne.datasets import eegbci
from mne.io import concatenate_raws, read_raw_edf
from gedai import Gedai
from gedai.viz import plot_mne_style_overlay_interactive
subjects = [1] # may vary
runs = [4, 8, 12] # may vary
raw_fnames = eegbci.load_data(subjects, runs, update_path=True)
raws = [read_raw_edf(f, preload=True) for f in raw_fnames]
# Concatenate runs from the same subject
raw = concatenate_raws(raws)
# Make channel names follow standard conventions
eegbci.standardize(raw)
# Crop to the first 15 seconds for demonstration purposes
# (Remove or adjust this for full data analysis)
raw.crop(0, 15)
raw.pick("eeg").load_data().apply_proj()
# Apply average reference (standard preprocessing for EEG)
raw.set_eeg_reference("average", projection=False)
Extracting EDF parameters from /home/runner/mne_data/MNE-eegbci-data/files/eegmmidb/1.0.0/S001/S001R04.edf...
Setting channel info structure...
Creating raw.info structure...
Reading 0 ... 19999 = 0.000 ... 124.994 secs...
Extracting EDF parameters from /home/runner/mne_data/MNE-eegbci-data/files/eegmmidb/1.0.0/S001/S001R08.edf...
Setting channel info structure...
Creating raw.info structure...
Reading 0 ... 19999 = 0.000 ... 124.994 secs...
Extracting EDF parameters from /home/runner/mne_data/MNE-eegbci-data/files/eegmmidb/1.0.0/S001/S001R12.edf...
Setting channel info structure...
Creating raw.info structure...
Reading 0 ... 19999 = 0.000 ... 124.994 secs...
No projector specified for this dataset. Please consider the method self.add_proj.
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
GEDAI🔗
To use spectral GEDAI, we initialize the Gedai
object by specifying the wavelet_levels parameter, which defines the
number of frequency bands to decompose the EEG data into. Each level
corresponds to a specific frequency band, allowing for targeted denoising
within those bands. It is also possible to define the type of wavelet used
for the decomposition by setting the wavelet_type parameter.
gedai = Gedai(wavelet_type="haar", wavelet_level=5)
Model Fitting🔗
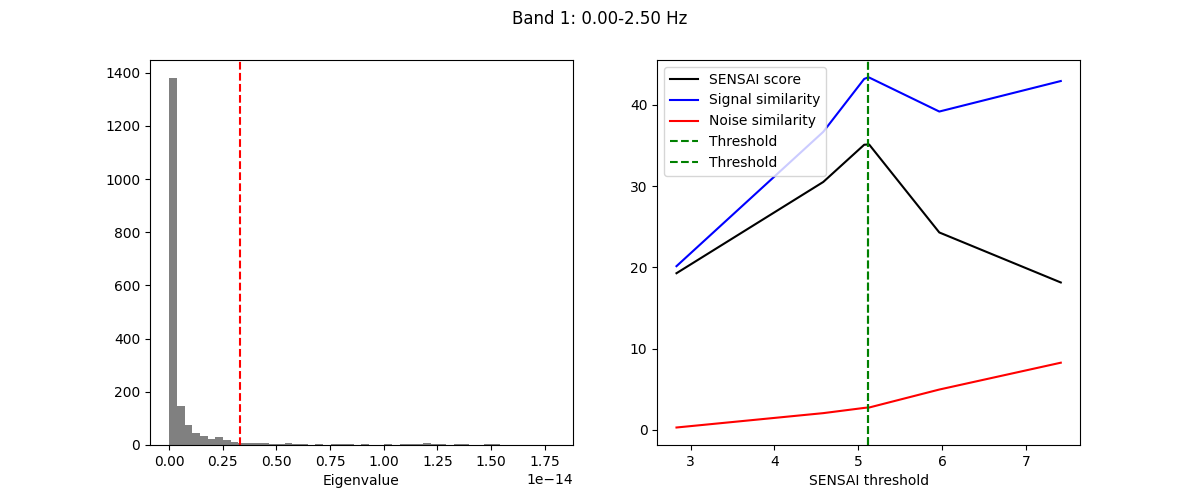
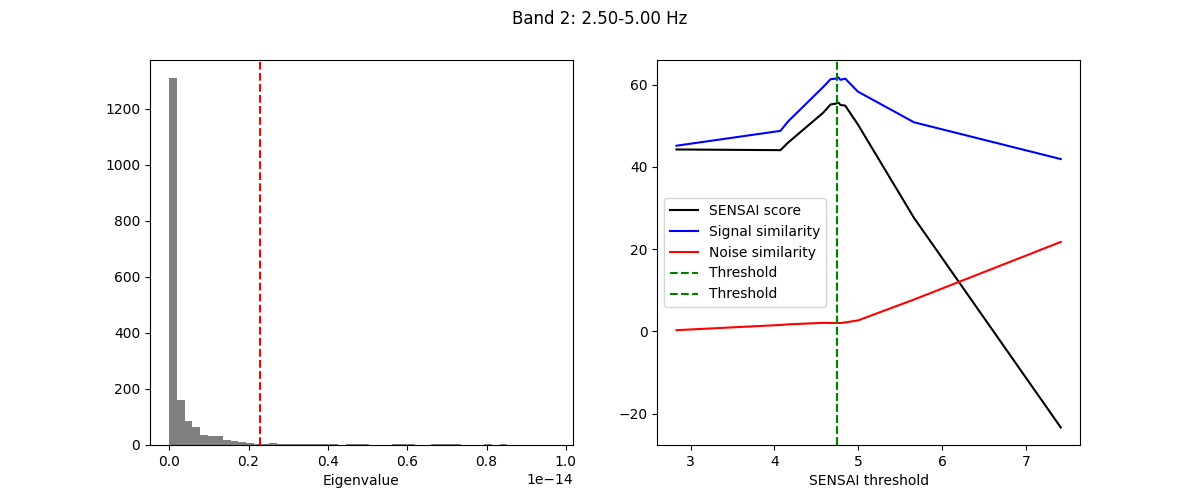
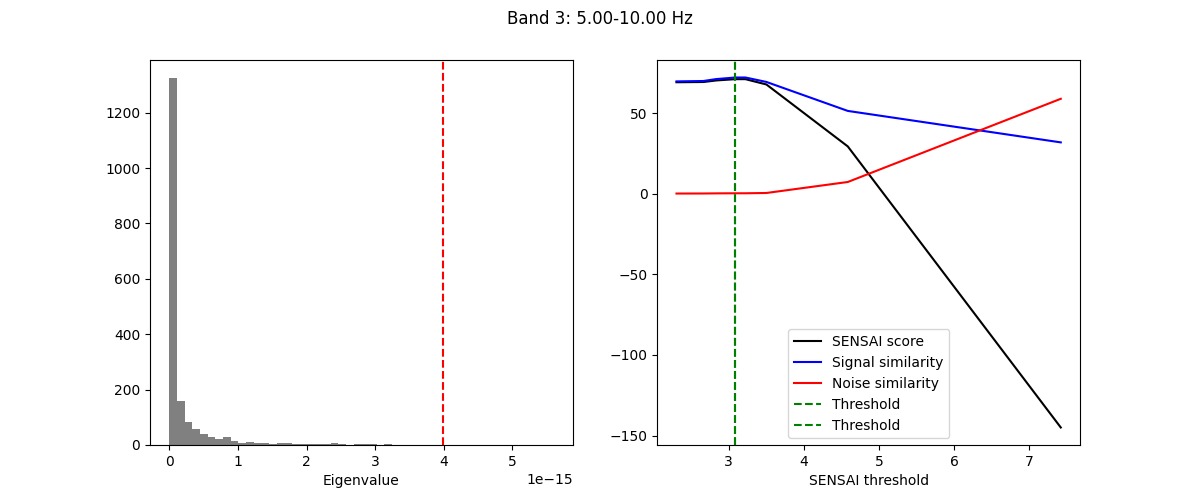
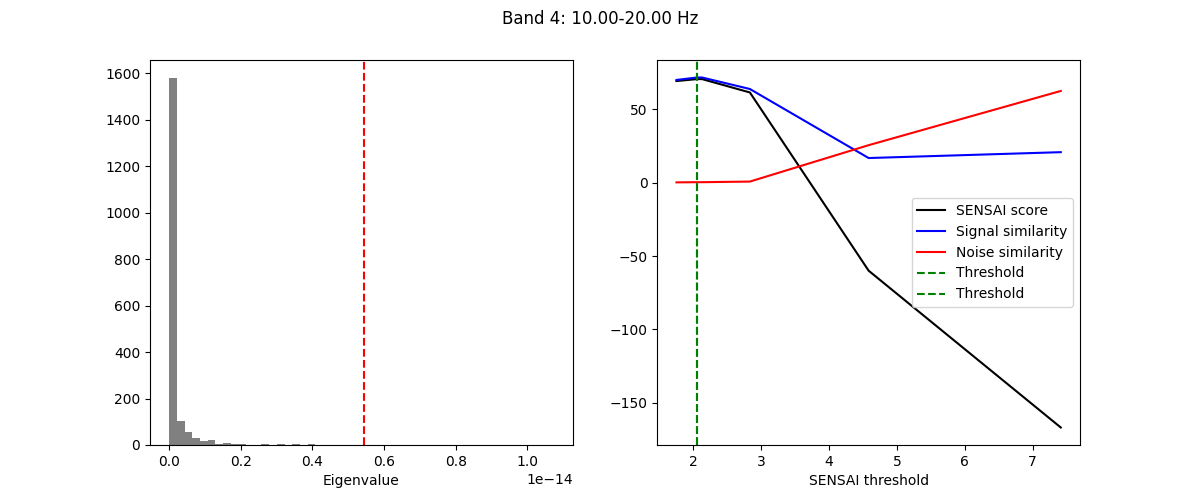
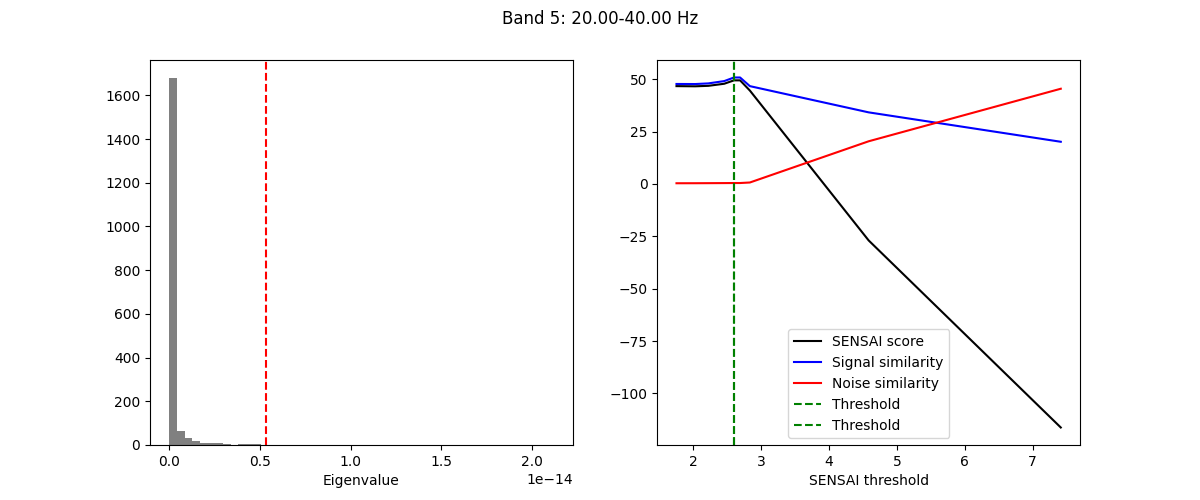
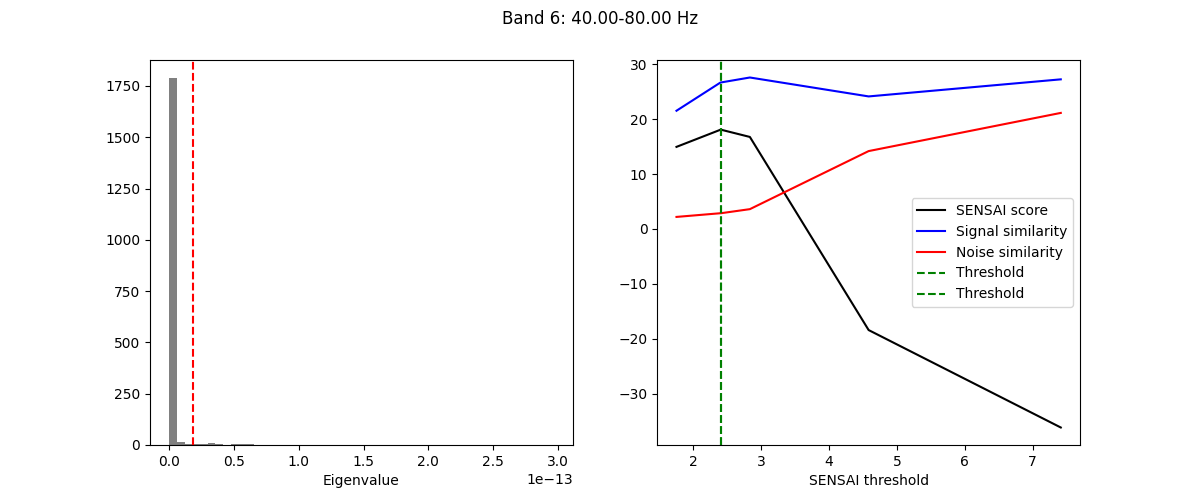
The fitting process of spectral GEDAI is similar to that of the standard
GEDAI. For each wavelet level (i.e., frequency band), the fitting process
estimates the optimal threshold to distinguish between signal and noise
components.
gedai.fit_raw(raw, verbose=True)
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
Note
Since spectral GEDAI uses spectral decomposition, the fitting
process will automatically adjust the epoch duration to ensure that
each epoch contains a number of samples appropriate for the wavelet
decomposition.
Transform the Data (Denoising)🔗
Once fitted, the Spectral GEDAI model can be used to remove artifacts
and noise from the data. The transform operation projects out the noise
components while preserving the brain signals for each frequency band
separately before recombining them.
raw_corrected = gedai.transform_raw(raw, verbose=False)
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
Warning
Since the spectral GEDAI operates on epoched data internally,
some frequency content more particularly in lower frequency bands may
be not be captured properly if the epoch duration is too short. On the
other hand, using very long epochs may prevent to capture short
transient artifacts. Setting the wavelet_low_cutoff parameter to a
value of the order of 1 / epoch_duration can help mitigate this
issue by excluding lower frequency bands that may not be well
estimated during the fitting process.
plot_mne_style_overlay_interactive(raw, raw_corrected)
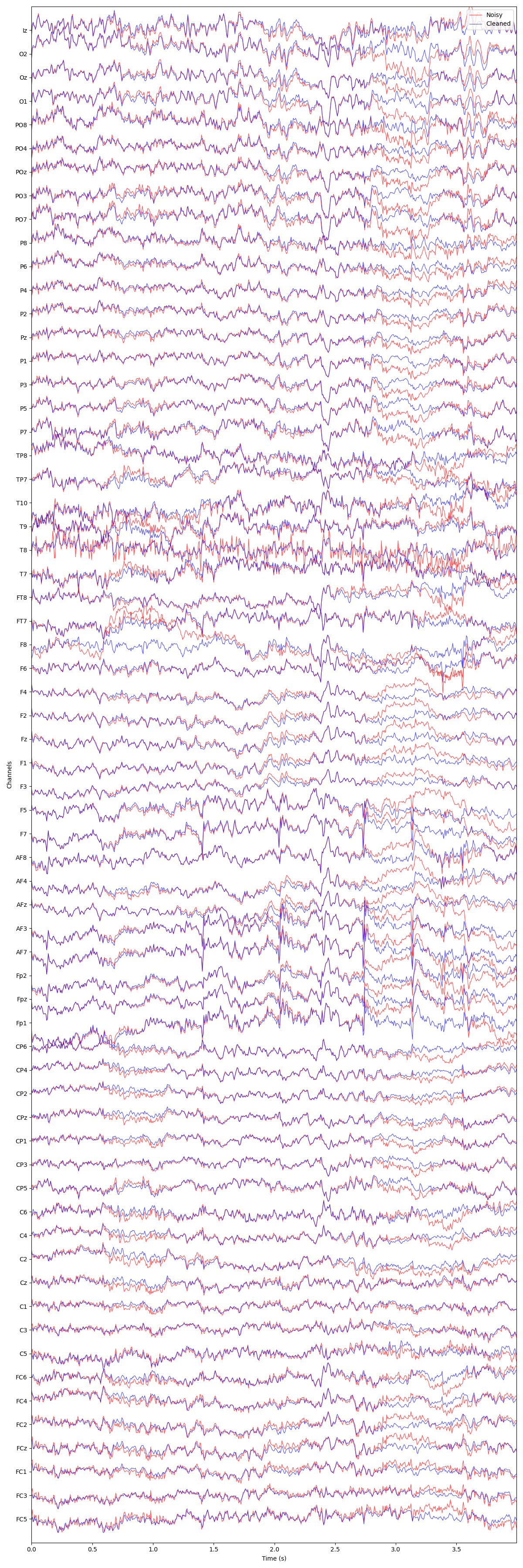
(<Figure size 1200x3600 with 1 Axes>, <Axes: xlabel='Time (s)', ylabel='Channels'>)
Recommended pipeline🔗
For optimal results, we recommend to first fit the standard GEDAI on
broadband data with a conservative noise_multiplier (e.g., 6.0) to
preserve most neural signals while only removing large artifacts. Then, use
the resulting cleaned data to fit the spectral GEDAI model. This two-step
approach leverages the strengths of both methods, ensuring effective artifact
removal while maintaining the integrity of neural signals across different
frequency bands.
gedai_broadband = Gedai()
gedai_broadband.fit_raw(raw, noise_multiplier=6.0)
raw_broadband_corrected = gedai_broadband.transform_raw(raw, verbose=False)
gedai_spectral = Gedai(wavelet_type="haar", wavelet_level=5, wavelet_low_cutoff=2)
gedai_spectral.fit_raw(raw_broadband_corrected, noise_multiplier=3.0)
raw_spectral_corrected = gedai_spectral.transform_raw(
raw_broadband_corrected, verbose=False
)
plot_mne_style_overlay_interactive(raw, raw_spectral_corrected)
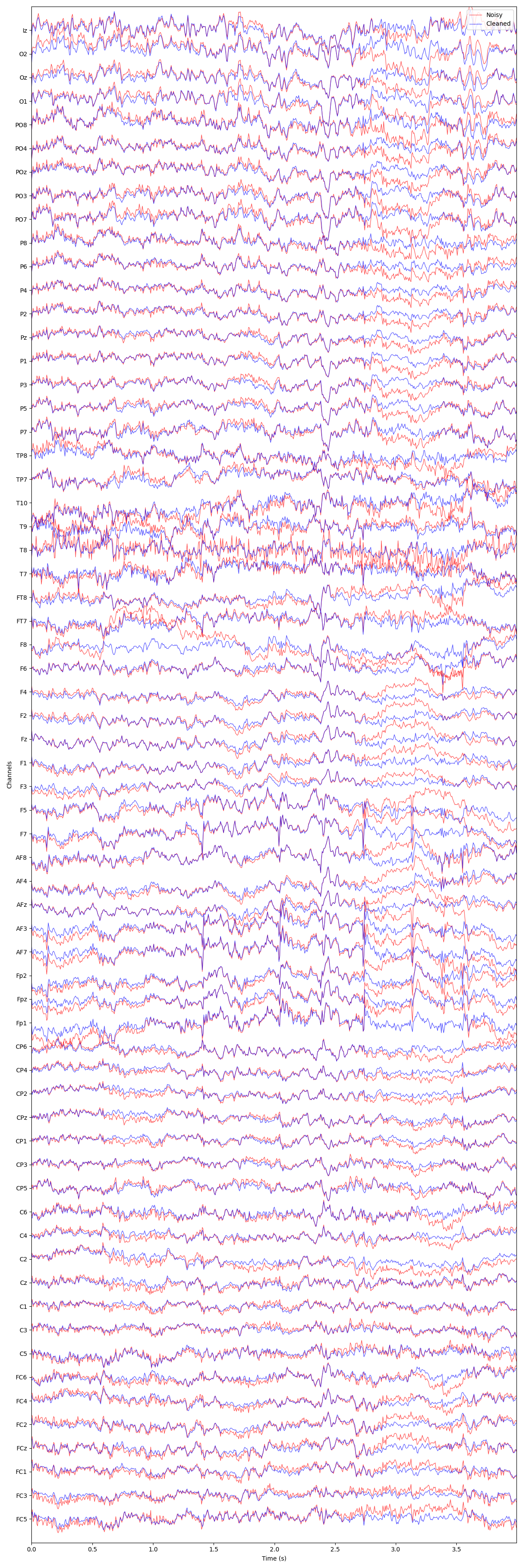
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
EEG channel type selected for re-referencing
Applying average reference.
Applying a custom ('EEG',) reference.
(<Figure size 1200x3600 with 1 Axes>, <Axes: xlabel='Time (s)', ylabel='Channels'>)
Total running time of the script: (12 minutes 1.776 seconds)
Estimated memory usage: 284 MB